A Model to Estimate SARS-CoV-2-Positive Americans
 For several reasons, testing for Covid-19 in the United States is deeply inadequate, which means that the true number of people infected with the SARS-CoV-2 virus is necessarily much higher than the reported, or confirmed, cases. But how much higher is the true number of “positives?” Having a good sense of this number is critical for planning, policy, communication, and mitigation.
For several reasons, testing for Covid-19 in the United States is deeply inadequate, which means that the true number of people infected with the SARS-CoV-2 virus is necessarily much higher than the reported, or confirmed, cases. But how much higher is the true number of “positives?” Having a good sense of this number is critical for planning, policy, communication, and mitigation.
In “Estimating SARS-CoV-2-positive Americans using deaths-only data,” released as a pre-print today on arXiv, our colleagues James Johndrow, Kristian Lum, and Patrick Ball built a model for estimating the true number of positives, using what they believe is the most reliable available data—number of deaths.
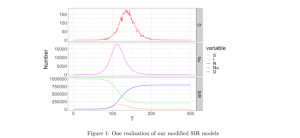
The case count data is highly unreliable because it depends on testing. How many tests are available varies in different places, and the rules that doctors and public health authorities apply for who gets tested changes rapidly. The number of deaths attributable to Covid-19 is likely to be relatively more reliable because it’s much easier to observe a death than a (possibly asymptomatic) infection. Johndrow, Lum, and Ball use a Bayesian model that extends the standard Susceptible-Infectious-Removed (SIR) model. They use data from clinical studies about the progression of the disease (from infection to death or recovery) and existing estimates of the R0 (the degree of infectiousness). Because they do not use case reports about the number of infections, they work backwards from the time series of observed deaths to the process of infection. They write in their introduction:
The goal of this work is not to provide specific forecasts of the infected population and likely deaths, although our initial estimates here fit the observed patterns of deaths. Accurate forecasts will require location-specific measures of containment and treatment efficacy, as well as age- and comorbidity-specific infected fatality rates. We don’t have these data at present, but our model could incorporate them as better information becomes available. Our paper offers a modeling approach using minimal but probably-good data, describes a likelihood and priors, is fitted to data, and is underpinned by a widely-used epidemiological model that is designed to approximate the real dynamics of disease spread.
What we think is most valuable and perhaps novel about this model is that the authors use the most reliable available data, and they can estimate a meaningful error, the amount by which the model is likely wrong. They use a well-understood and deeply researched theory from epidemiology—SIR—that is appropriate to this situation. The result is a model that’s principled, relatively simple, and extensible.
As HRDAG’s executive director, I am proud of this rapid work from our colleagues. HRDAG’s specialty is using statistical methods to draw defensible conclusions from incomplete, imperfect data. As this crisis continues to unfold, we will combine these skills with our collective prior experiences in public health (me), infectious disease modeling (Lum), and genetic phylogeny (Johndrow) to generate and illuminate good science in our attempt to answer to these crucial questions.
Acknowledgments
Support for this project was provided by grants from the John D. and Catherine T. MacArthur Foundation and the Oak Foundation. (For more information about HRDAG’s supporters, please see our Funding page.)
Find more articles about HRDAG research and resources regarding Covid-19.
